 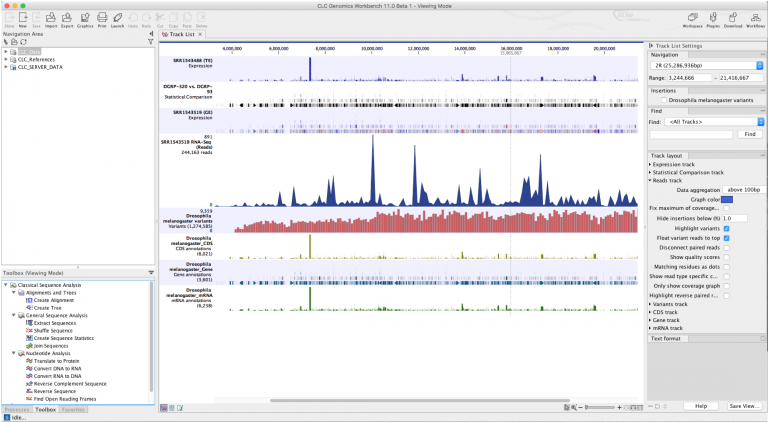
SNPs and genotypes were called using three different algorithms. We sequenced ~1% of the orangutan genome with 41-fold median coverage in 31 wild-born individuals from two populations. Our modifications facilitate generation of single-sample libraries, enabling individual genotype assignments instead of pooled-sample analysis. We present an improved RRL (iRRL) protocol that maximizes the generation of homologous DNA sequences, thus achieving improved genotyping-by-sequencing efficiency. From the bioinformatical perspective, the reliance of most studies on a single SNP caller disregards the possibility that different algorithms may produce disparate SNP datasets. In the laboratory, current protocols require improvements with regards to sequencing homologous fragments to reduce the number of missing genotypes. Yet, generating such datasets remains challenging due to laboratory and bioinformatical issues. Like similar approaches, RRL sequencing reduces ascertainment bias due to simultaneous discovery and genotyping of single-nucleotide polymorphisms (SNPs) and does not require reference genomes. investigating only parts of the genome, is reduced-representation library (RRL) sequencing. One approach to reduce genome complexity, i.e. It's only 10 or so pages and it will give you a better idea of what you can expect regarding your own situation.High-throughput sequencing has opened up exciting possibilities in population and conservation genetics by enabling the assessment of genetic variation at genome-wide scales. I hope this helps! I recommend reading the actual paper. MIRA (3 days) was not distinguished, barely doing better than SeqMan at anything yet taking by far the longest.CAP3 (1 day) was not distinguished, doing no better than Newbler 2.5 or SeqMan at anything yet taking longer.SeqMan is also fantastic, generating the most unique sequences and the largest assembly but taking much longer than Newbler (6 hrs).Newbler 2.5 is fantastic overall, generating an overall very large assembly in a moderate amount of time (45 min).CLC is the fastest by far (4 min) and gathers a lot of unique contigs due to the de Bruijn graph algorithm used BUT is inaccurate and generates an overall smaller assembly.Newbler 2.3 is the worst and shouldn't be used.It compares Newbler 2.3, Newbler 2.5, CAP3, CLC, SeqMan, and MIRA. 
If anyone is still interested in this subject, I found this paper to be extremely helpful:Ĭomparing de novo assemblers for 454 transcriptome data (don't hesitate to share your ideas on mira3 or other open software also :) As you mention, the scary (at first) cost of the license compares lightly to the total cost of one (not mentioning that you will probably use it for many) next gen projects. Overall, our experience with CLC HAS been very satisfactory and we will likely continue to use it in the near future. The steps involved in the alignment lead to more gene chimeras and strange coverage patterns within each contig that may be expected from a 'correct' approach. The reason I am looking into mira3 now is that, it appears that the de novo alignment algorithm is not totally appropriate for RNA-seq projects. It is, however, highly suggested to use the software on a 64 bit system with plenty of ram (8 gigs and up).Īll this comes at the price of some flexibility and some transparency, I guess. This we have done repeatedly and new comers in the lab get quickly to their results without too much of a chock. sff or fasta/fasta.qual data, trim sequences according to different criteria, do a de novo assembly, save consensus sequences from the contigs, do a reference assembly (possibly with only a subset of sequences), look for SNPs, export SNP tables and ACE assembly files. For example, using menus, it is easy to import. I would say that the main strength of CLC is it's easiness of use, mainly for non-computer-oriented biologists, or even for non-hardcore-linux users. We have mostly used the software to toy with RNA-seq 454 data in non-model species, so our expertise is with de-novo assembly of expressed sequences, namely cDNA from RNA containing poly-A tails. 
Nothing that you couldn't find in the open source world, but well put together. The software IS really a well integrated resources with a lot of small functionalities. We have been using CLC Genomic Workbench for the last year with great results. I work in a lab mainly into evolution and genomics in fishes. I am presently going the opposite way from what you do, coming from CLC, I am exploring (anew) the possibilities of open software (mainly mira3 now) for 454 assembly. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
